Shear & B.O. Dodge (1927)
What is Neurospora crassa?
Neurospora crassa (Fig. 1) is colloquially known as the red bread mold, as it was initially discovered forming reddish or orange colonies on bread and pastries in French bakeries in 1843. It is a filamentous fungus that belongs to the phylum Ascomycota. Despite its initial discovery as a food contaminant, it was concluded that it is neither toxic nor pathogenic, neither to humans nor to animals and plants. There are no known cases of home contaminations or diseases caused by this fungus (1, 2, 3).
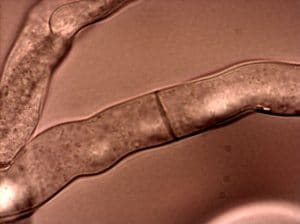
How is Neurospora crassa related to science?
Being nonpathogenic and easy to grow (minimal nutrient requirements, fast growth, and a short yet complex life cycle), this fungus was chosen as a model organism for biochemical research by Beadle and Tatum in 1941. Their research led to the “one gene-one enzyme” hypothesis, which proposed that every gene within the genome of a living organism was in charge of the synthesis and function of a single enzyme. This protein catalyzes the speed of a chemical reaction in the cell. Until this hypothesis, biochemistry and genetics were separate fields of research, both still in the early stages of scientific discovery. This discovery led to the creation of a new field, biochemical genetics, and emphasized the importance of these research fields in understanding how living organisms function (1, 2).

Even though this hypothesis evolved and was changed into “one gene-one protein/polypeptide” throughout the years, with the realization that not every protein in the cell has the catalytic role of an enzyme, the discovery earned N. crassa a very important place in the history of science (1,2). Since then, it has been used as a model organism for multiple other research topics, such as amino acid synthesis, DNA and RNA studies, sugar metabolism, circadian rhythms, cell differentiation, and others (1, 4).
Is Neurospora crassa dangerous?
Before its inclusion in scientific research, N. crassa was known as a common food contaminant in bakeries. At that time, Neurospora species were classified under a different scientific name, in the genus Motilia. The species most commonly isolated in bakeries was Motilia sitophila, the asexual or imperfect form of Neurospora sitophila (5, 6). Molds whose sexual states have not yet been discovered or do not have a sexual state in their life cycles are sometimes referred to as fungi imperfecti. These molds were often thought to belong to different genera before discovering their sexual states, hence the classification of Neurospora in the genus Motilia.
Neurospora species were often found in bakeries, contaminating both flour and baked goods. They were quite difficult to eradicate and caused huge financial losses until the introduction of fungal growth inhibitors, along with improved hygiene (4, 5, 6). Nowadays, N. crassa can more commonly be found growing on sugar cane bagasse (5), as a saprophyte in the soil or on tree bark, especially on burned trees after a forest fire, since the germination of its spores is triggered by thermal shock (4, 10).
How does Neurospora crassa reproduce?
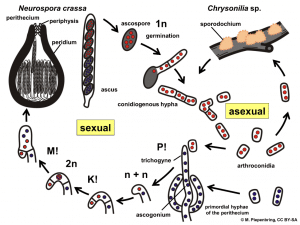
Neurospora crassa, being a member of Ascomycota, has a complex life cycle. Ascomycota are named so because their spores are formed in a sack-like structure (“ascus” is Latin for a sack or a bag). Some of these fungi can have both sexual and asexual forms of reproduction (Fig. 2). When the environmental conditions aren’t ideal for sexual reproduction or in the absence of individuals with the opposite mating type, these fungi will reproduce asexually by a simple fragmentation of their mycelium or through conidia (asexual spores). Once the hyphal fragments or the conidia are dispersed, new hyphae will form from them. However, as there was no exchange and recombination of genetic information with another member of the same species, the genetic material of the new colony is identical to the original mycelium, meaning it is a clone (4, 7, 8). In these situations, the main source of genetic diversity is genetic mutations during the replication of DNA.
Fungi don’t have male and female members of the species in the classic sense. Instead, they have mating types that form different sexual structures. In the case of N. crassa, they are referred to as mating types A and a. For sexual reproduction to happen, the primary mycelia of the opposite mating type individuals need to come in contact. One of the mating types forms a structure called a perithecium. The perithecium is made out of two parts: the ascogonium and the trichogyne. The trichogyne is a slender hypha that extends toward the conidia of the other mating type until the cells connect and fuse. New hyphae form from the ascogonium called ascogenous hyphae.
The top of the ascogenous hyphae forms a hook-like shape, which is divided into three cells. The top and the bottom cell have only one nucleus, while the middle cell contains both nuclei from the two mating types (n + n). The two nuclei fuse, forming a zygote, a “fertilized” cell. It has two sets of chromosomes (2n) that go through recombination, the process of rearranging genetic information between the chromosomes. The zygote then splits back into cells with a single set of chromosomes (n). The result of the divisions is an ascus containing 8 ascospores. A heat shock often activates the germination of ascospores. Once these spores germinate, they form a new mycelium, with a brand new combination of genes (7 – 10).
Neurospora crassa genome
The genome of N. crassa was fully sequenced in 2003. It was shown that this mold has around 10 000 different genes that all contain information for the synthesis of a protein (protein-coding genes), some of which share with other eukaryotic organisms and some specific only to N. crassa. Compared to other commonly used model organisms, N. crassa has almost twice as many protein-coding genes as Saccharomyces cerevisiae and almost as many genes as the fruit fly, Drosophila melanogaster (2, 11).
Neurospora crassa circadian rhythm
The name “circadian rhythm” is derived from the Latin words “circa”, which means “approximately”, and “diem”, which means “day”. Thus, a circadian rhythm represents a physiological process in an organism’s body (be it an animal, plant, or microorganism) that occurs and fluctuates in the same manner roughly every 24 hours and is influenced by light, temperature, and other external factors. The most well-known circadian rhythm is our sleep-wake cycle, which depends on the amount of light we are exposed to (whether it is night or day) and the synthesis of the hormone melatonin (12).
In N. crassa, a circadian rhythm has been linked to its asexual reproduction. Neurospora crassa not only differentiates between night and day, based on light exposure, but it also adjusts the synthesis of its asexual spores (conidia) to the time of day. This was tested by growing mycelia in glass tubes, called “race tubes”. Depending on the time of day, the fungus would switch between mycelium growth and the development of aerial hyphae and conidia. This process occurs on a basic cellular level through the expression of a gene called the frequency gene (frq). (1, 13)
Neurospora crassa uses in agriculture
As previously stated, N. crassa is safe for people to interact with and has even been used as food by certain indigenous tribes in Java, Borneo, and Brazil. The fungus is either collected as a direct food source, as is the case of Iban people in Borneo gathering it from burnt rice fields, or used to process other food. For example, it is used in Brazil to create fermented drinks through cassava processing (3).
Neurospora crassa has shown great potential in biomass processing, as it is highly effective in breaking down components of the plant cell wall and plant biomass – cellulose and hemicellulose. It synthesizes important enzymes that break down these substances into simple sugars, like xylose and arabinose, which can then be used for fermentation or the production of liquid fuels (14, 15).

Did you know?
The #1 toxic mold type found in kitchens in Canada is the Penicillium/Aspergillus mold group?! Find out more exciting mold stats and facts on our mold statistics page.
References
- Roche, C. M., Loros, J. J., McCluskey, K., & Glass, N. L. (2014). Neurospora crassa: looking back and looking forward at a model microbe. American journal of botany, 101(12), 2022-2035.
- Arnold, J., & Hilton, N. (2003). Revelations from a bread mould. Nature, 422(6934), 821-822.
- Perkins, D. D., & Davis, R. H. (2000). Evidence for safety of Neurospora species for academic and commercial uses. Applied and environmental microbiology, 66(12), 5107-5109.
- King, R. C. (Ed.). (2013). Handbook of genetics: volume 1 bacteria, bacteriophages, and fungi(Vol. 1). Springer Science & Business Media.
- Shear, C. L., & Dodge, B. O. (1927). Life histories and heterothallism of the red bread-mold fungi of the Monilia sitophila group(pp. 1019-1042). Washington, DC: US Government Printing Office.
- Yassin, S., & Wheals, A. (1992). Neurospora species in bakeries. Journal of applied bacteriology, 72(5), 377-380.
- Seale, T. (1973). Life cycle of Neurospora crassa viewed by scanning electron microscopy. Journal of bacteriology, 113(2), 1015-1025.
- Wallen, R. M., & Perlin, M. H. (2018). An overview of the function and maintenance of sexual reproduction in dikaryotic fungi. Frontiers in microbiology, 9, 503.
- Vukojević, J. (2012). Praktikum iz mikologije i lihenologije. NNK Internacional
- Aramayo, R., & Selker, E. U. (2013). Neurospora crassa, a model system for epigenetics research. Cold Spring Harbor Perspectives in Biology, 5(10), a017921.
- Galagan, J. E., Calvo, S. E., Borkovich, K. A., Selker, E. U., Read, N. D., Jaffe, D., … & Birren, B. (2003). The genome sequence of the filamentous fungus Neurospora crassa. Nature, 422(6934), 859-868.
- Brown, G. M. (1994). Light, melatonin and the sleep-wake cycle. Journal of Psychiatry and Neuroscience, 19(5), 345.
- Baker, C. L., Loros, J. J., & Dunlap, J. C. (2012). The circadian clock of Neurospora crassa. FEMS microbiology reviews, 36(1), 95-110.
- Glass, N. L., Schmoll, M., Cate, J. H., & Coradetti, S. (2013). Plant cell wall deconstruction by ascomycete fungi. Annual review of microbiology, 67, 477-498.
- Tian, C., Beeson, W. T., Iavarone, A. T., Sun, J., Marletta, M. A., Cate, J. H., & Glass, N. L. (2009). Systems analysis of plant cell wall degradation by the model filamentous fungus Neurospora crassa. Proceedings of the National Academy of Sciences, 106(52), 22157-22162.

Get Special Gift: Industry-Standard Mold Removal Guidelines
Download the industry-standard guidelines that Mold Busters use in their own mold removal services, including news, tips and special offers:

Written by:
Ema Zivkovic
Mycologist
Mold Busters
Edited by:
Dusan Sadikovic
Mycologist – MSc, PhD
Mold Busters
